Z-filtered MQMAS
AIM: We provide Mathematica-5 notebooks and SIMPSON1.1.1 Tcl scripts to optimize the echo and antiecho amplitudes for z-filtered MQMAS NMR experiment applied to half-integer quadrupole spin.
J.-P. Amoureux and coworkers [Z filtering in MQ MAS NMR, J. Magn. Reson. A 123, 116-118 (1996)] present the z-filter approach which allows the symmetrization of both echo and antiecho transfer pathways, generating a 2D pure absorption MQMAS spectrum.
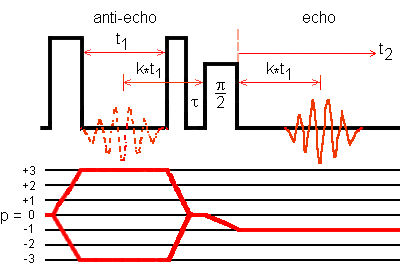
Fig. 1: Z-filtered MQMAS pulse sequence and coherence transfer
pathways for 3Q echo and -3Q antiecho of a spin I = 5/2 system.
0Q -> 3Q -> 0Q -> -1Q is the 3Q echo transfer pathway.
0Q -> -3Q -> 0Q -> -1Q is the -3Q antiecho transfer pathway.
The echo amplitude and the antiecho amplitude have the same sign.
Method: We simulate the echo and the antiecho amplitudes of a spin I = 5/2 with increasing second-pulse duration in a powder rotating at the magic angle, using Mathematica-5 notebooks and SIMPSON1.1.1 Tcl scripts.
The parameters for these simulations are:
- Nucleus: 27Al
- Spin: 5/2
- 27Al Larmor frequency: 208.61889974 MHz
- Proton Larmor frequency: 800 MHz
- Only 3Q and -3Q coherences belonging to the second diagonal of the density matrix are taken into account for the simulation
- Amplitude of the strong radio-frequency pulse: 90 kHz
- Amplitude of the weak radio-frequency pulse: 9.3 kHz
- First-pulse duration: 4 μs
- Initial duration of the second pulse: 0
- Final duration of the second pulse: 4 μs
- Pulse duration increment: 0.25 μs
- Number of the second-pulse duration increment: 17
- Third-pulse duration: 9 μs
- Rotor spinning speed: 5 kHz
- Quadrupole interaction: first and second orders
- Quadrupole coupling constant: 5 MHz
- Asymmetry parameter: -1 for notebook and 1 for Tcl script
- Crystal file: rep100_simp for notebook and rep100.cry for Tcl script
- Number of summation steps of the Euler angle γ of the rotor: 10
(A) Mathematica-5 notebook
(1) Preliminary
| Optimization with |
Notebook |
|---|---|
| strong pulse p1 | zfilter_P1.nb (pdf) |
| strong pulse p2 | zfilter_P2.nb (pdf) |
| soft pulse p3 | zfilter_P3.nb (pdf) |
- Download Mathematica-5 notebook zfilter_P2.nb, that for MAS NMR utilities QUADRUPOLE_1_0.nb (the corresponding PDF file), and the crystal file rep100_simp.
- Save these three files into the software Mathematica-5 folder. Forbidden the Operating System of your computer to include extra file extension to rep100_simp by providing the file name with double quotes such as "rep100_simp".
- Open QUADRUPOLE_1_0.nb file with Mathematica-5.
- Press "Ctrl-A" to select the notebook, then press "Shift-enter" to start the notebook. (Some warning messages appear but they have no consequences on the results.) A new file called QUADRUPOLE is created in Mathematica-5 folder.
(2) Simulation
- Open zfilter_P2.nb file with Mathematica-5.
- Press "Ctrl-A" to select the notebook, then press "Shift-enter" to start simulation. (Some warning messages precede the simulation.) At the end a data file called zfilter_P2 is created in Mathematica-5 folder. MS Excel can open this data file for graphic representation.
(B) SIMPSON1.1.1 Tcl script
(1) Preliminary
| Optimization with | SIMPSON1.1.1 |
|---|---|
| strong pulse p1 | zfilter_p1.in |
| strong pulse p2 | zfilter_p2.in |
| soft pulse p3 | zfilter_p3.in |
Download and save zfilter_p2.in file into the software SIMPSON1.1.1 folder.
(2) Simulation
- Run zfilter_p2.in in a DOS window.
- The simulated signal amplitudes are saved in the file called zfilter_p2.fid in SIMPSON1.1.1 folder. MS Excel also can open this data file for graphic representation.
(C) Result
Figure 2 represents the simulated data. Notebook and Tcl script provide the same curve.
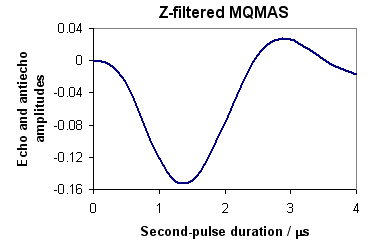
Fig. 2: 27Al z-filtered 3Q echo and -3Q antiecho amplitudes versus the second-pulse duration.
We also provide notebooks and SIMPSON1.1.1 Tcl scripts where all the coherences belonging to the same MQ coherence transfer pathway are considered:
| Optimization with |
Notebook | SIMPSON 1.1.1 |
|---|---|---|
| strong pulse p1 | zfilter_P1S.nb, (pdf) |
zfilter_p1S.in |
| strong pulse p2 | zfilter_P2S.nb, (pdf) |
zfilter_p2S.in |
| soft pulse p3 | zfilter_P3S.nb, (pdf) |
zfilter_p3S.in |
These files allow us to compare simulated data with those obtained with SPAM MQMAS approach.
Nicolas Malicki and coworkers [Multiplex MQMAS NMR of quadrupolar nuclei, Solid State Nucl. Magn. Reson. 28, 13-21 (2005)] present the Multiplex z-filter approach which reduces the experimental time considerably.
